Overview of Estimation of Distribution Algorithms (EDAs)
Abstract
This work shows how Estimation of Distribution Algorithms (EDAs) can be applied to combinatorial optimization problems. Two approximation to the problem are introduced: one for the discrete EDAs and other for continuous ones. The EDA approach is illustrated with the Griewangk problem.
1 Introduction
Estimation of Distribution Algorithms (EDAs) [ Mühlenbein and Paaß1996, Larrañaga and Lozano2001] constitute a new and promising stochastic heuristic for optimization that can be considered as member of the family of evolutionary computation heuristics. Due to their recent development, their application to classic and real optimization problems is still starting. This work presents a first approximation to a type of combinatorial optimization problems -searching for the best permutation- that have a big relevance in areas such as Operational Research and Machine Learning.
2 Estimation of Distribution Algorithms
2.1 Introduction
EDAs [ Mühlenbein and Paaß1996, Larrañaga and Lozano2001] are non-deterministic, stochastic heuristic search strategies that form part of the evolutionary computation approaches, where number of solutions or individuals are created every generation, evolving once and again until a satisfactory solution is achieved. In brief, the characteristic that most differentiates EDAs from other evolutionary search strategies such as GAs is that the evolution from a generation to the next one is done by estimating the probability distribution of the fittest individuals, and afterwards by sampling the induced model. This avoids the use of crossing or mutation operators, and the number of parameters that EDAs require is reduced considerably.
|
In EDAs, individuals are not said to contain genes, but variables which dependencies have to be analyzed. Also, while in other heuristics from evolutionary computation the interrelations between the different variables representing the individuals are kept in mind implicitly (e.g. building block hypothesis), in EDAs the interrelations are expressed explicitly through the joint probability distribution associated with the individuals selected at each iteration. The task of estimating the joint probability distribution associated with the database of the selected individuals from the previous generation constitutes the hardest work to perform, as this requires the adaptation of methods to learn models from data developed in the domain of probabilistic graphical models.
Figure 1 shows the pseudocode of EDA, in which we distinguish four main steps in this approach:
- At the beginning, the first population D0 of M individuals is generated, usually by assuming an uniform distribution (either discrete or continuous) on each variable, and evaluating each of the individuals.
- Secondly, a number N (N £ M) of individuals are selected, usually the fittest.
- Thirdly, the n-dimensional probabilistic model that better expresses the interdependencies between the n variables is induced.
- Next, the new population of M new individuals is obtained by simulating the probability distribution learnt in the previous step.
Next, some notation is introduced. Let X = (X1, ¼, Xn) be a set of random variables, and let xi be a value of Xi, the ith component of X. Let y = (xi)Xi Î Y be a value of Y Í X. Then, a probabilistic graphical model for X is a graphical factorization of the joint generalized probability density function, r(X = x) (or simply r(x)). The representation of this model is given by two components: a structure and a set of local generalized probability densities.
Regarding the structure of the model, the structure S for X is a directed acyclic graph (DAG) that describes a set of conditional independences between the variables on X. PaiS represents the set of parents -variables from which an arrow is coming out in S- of the variable Xi in the probabilistic graphical model, the structure of which is given by S. The structure S for X assumes that Xi and its non descendents are independent given PaiS, i = 2, ¼, n. Therefore, the factorization can be written as follows:
|
(1) |
Furthermore, regarding the local generalized probability densities associated with the probabilistic graphical model, these are precisely the ones appearing in Equation 1.
A representation of the models of the characteristics described above assumes that the local generalized probability densities depend on a finite set of parameters qS Î QS, and as a result the previous equation can be rewritten as follows:
|
(2) |
After having defined both components of the probabilistic graphical model, the model itself will be represented by MO = (S,qS).
2.2 EDAs in discrete domains
In the particular case where every variable Xi Î X is discrete, the probabilistic graphical model is called Bayesian network. If the variable Xi has ri possible values, xi1, ¼, xiri, the local distribution, p (xi | paij,S,qi) is:
|
(3) |
All the EDAs are classified depending on the maximum number of dependencies between variables that they accept (maximum number of parents that any variable can have in the probabilistic graphical model).
2.2.1 Without interdependencies
The Univariate Marginal Distribution Algorithm (UMDA) [ Mühlenbein1998] is a representative example of this category, which can be written as:
|
(4) |
2.2.2 Pairwise dependencies
An example of this second category is the greedy algorithm called MIMIC (Mutual Information Maximization for Input Clustering) [ De Bonet et al. 1997]. The main idea in MIMIC is to describe the true mass joint probability as closely as possible by using only one univariate marginal probability and n - 1 pairwise conditional probability functions.
2.2.3 Multiple interdependencies
We will use EBNA (Estimation of Bayesian Network Algorithm) as an example of this category. The EBNA approach was introduced for the first time in [ Etxeberria and Larrañaga1999], where the authors use the Bayesian Information Criterion (BIC) as the score to evaluate the goodness of each structure found during the search.
The initial model MO0 in EBNA, is formed by its structure S0 which is an arc-less DAG and the local probability distributions given by the n unidimensional marginal probabilities p(Xi = xi)=[1/(ri)], i = 1, ¼, n -that is, MO0 assigns the same probability to all individuals. The model of the first generation -MO1- is learnt using Algorithm B [ Buntine91], while the rest of the models are learnt following a local search strategy that received the model of the previous generation as the initial structure.
2.2.4 Simulation in Bayesian networks
In EDAs, the simulation of Bayesian networks is used merely as a tool to generate new individuals for the next population based on the structure learned previously. The method used in this work is the Probabilistic Logic Sampling (PLS) [ Henrion88]. Following this method, the instantiations are done one variable at a time in a forward way, that is, a variable is not sampled until all its parents have already been so.
2.3 EDAs in continuous domains
In this section we introduce an example of the probabilistic graphical model paradigm that assumes the joint density function to be a multivariate Gaussian density.
The local density function for the ith variable is computed as the linear-regression model
|
(5) |
Local parameters are given by qi = (mi,bi, vi), where bi = (b1i, ¼, bi-1i)t is a column vector. Local parameters are as follows: mi is the unconditional mean of Xi, vi is the conditional variance of Xi given Pai, and bji is a linear coefficient that measures the strength of the relationship between Xj and Xi. A probabilistic graphical model built from these local density functions is known as a Gaussian network. Gaussian networks are of interest in continuous EDAs because the number of parameters needed to specify a multivariate Gaussian density is smaller.
Next, an analogous classification of continuous EDAs as for the discrete domain is done, in which these continuous EDAs are also classified depending on the number of dependencies they take into account.
2.3.1 Without dependencies
In this case, the joint density function is assumed to follow a n-dimensional normal distribution, and thus it is factorized as a product of n unidimensional and independent normal densities. Using the mathematical notation X º N (x;m, å), this assumption can be expressed as:
|
|||||||||||||||||||||||
2.3.2 Bivariate dependencies
An example of this category is MIMICcG [ Larrañaga et al. 2000], which is basically an adaptation of the MIMIC algorithm [ De Bonet et al. 1997] to the continuous domain.
2.3.3 Multiple dependencies
Algorithms in this section are approaches of EDAs for continuous domains in which there is no restriction in the learning of the density function every generation. An example of this category is EGNABGe (Estimation of Gaussian Network Algorithm) [ Larrañaga et al. 2000]. The method used to find the Gaussian network structure is a Bayesian score+search. In EGNABGe a local search is used to search for good structures.
2.3.4 Simulation of Gaussian networks
A general approach for sampling from multivariate normal distributions is known as the conditioning method, which generates instances of X by sampling X1, then X2 conditionaly to X1, and so on. The simulation of a univariate normal distribution can be done with a simple method based on the sum of 12 uniform variables.
3 Example of the Griewangk problem
This is a minimization problem proposed in [Törn and Zilinskas 1989]. The fitness function is as follows:
|
(2) |
The range of all the components of the individual is -600\leqslant xi \leqslant 600 , i = 1, ... , n, and the fittest individual corresponds to a value of 0, that only can be obtained when all the components of the individual are 0.
4 Brief description of the experiment
This section describes the experiments and the results obtained. Continuous EDAs were implemented in ANSI C++ language, and the ES techniques were obtained from Schwefel (1995). Some of the problems to optimize selected were not implemented in the ES program, and therefore some changes were made in order to include them in the ANSI C program.
The initial population for both continuous EDAs and ES was generated using the same random generation procedure based on a uniform distribution.
In EDAs, the following parameters were used: a population of 2000 individuals (M=2000), from which a subset of the best 1000 are selected (N=1000) to estimate the density function, and the elitist approach was chosen (that is, always the best individual is included for the next population and 1999 individuals are simulated).
The ES chosen was taken from Schwefel (1995). This is a standard (m + l)-strategy with discrete recombination in points of the search space and intermediary recombination for sigma vector. The value of the parameters was set to m = 15, l = 100, plus strategy, the ellipsoid is not able to rotate, recombination type for the x vector 2, recombination type for the sigma vector 3, lower bound to step sizes (absolute of 1E-30 and relative of 9.8E-07), parameter in convergence test (absolute of 1E-30 and relative of 9.8E-07).
For all the continuous EDAs the ending criterion was set to reach 301850 evaluations (i.e. number of individuals generated) or when the result obtained was closer than 1E-06 from the optimum solution to be found.
These experiments were all executed in a two processor Ultra 80 Sun computer under Solaris version 7 with 1 Gb of RAM.
4.1 Experimental results
The Griewangk problem is a very complex problem to optimize due to the many local minima it presents (see Figures 1 and 2). In the 10 dimension case all the mean values of the Table 1 the fitness values appear quite similar for all but the EMNAa algorithm. When performing the non parametric test for all the results we obtain though that differences are significant for both the fitness value and number of evaluations. The algorithm that shows the best behaviour is ES, closely followed by EGNABIC, EMNAglobal, EGNAee, MIMICc and UMDAc algorithms. When comparing ES and these five continuous EDAs we do not obtain statistically significant differences in the best fitness value (p = 0.101). On the other hand, the differences are significant in number of evaluations required for these algorithms (p < 0.001).
In the 50 dimension case the differences are bigger, and therefore the performance of the different algorithms can be seen more clearly. In this case ES arrived to the ending criterion and stopped the search without reaching a solution as well as the obtained with any of the EDAs. Differences among all the continuous EDAs are statistically significant looking at the results (p < 0.001), and the algorithms that quicker arrived to these results were EGNABGe, EGNAee, MIMICc and UMDAc, although the difference in fitness value of these was non significant (p = 0.505). The fact that EGNABIC did not converge as quick as the rest shows that this algorithm is more dependent on the complexity of the problem than the others.
Figure 1:Plot of the Griewangk funtion on its 600x600 scale. Function computed with scilab.
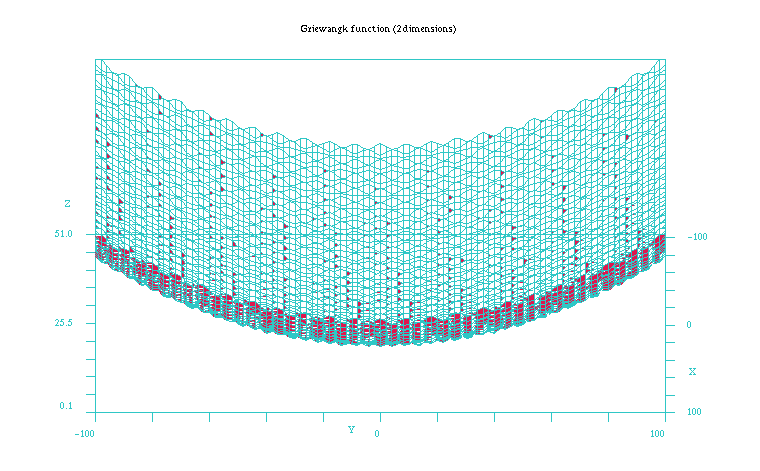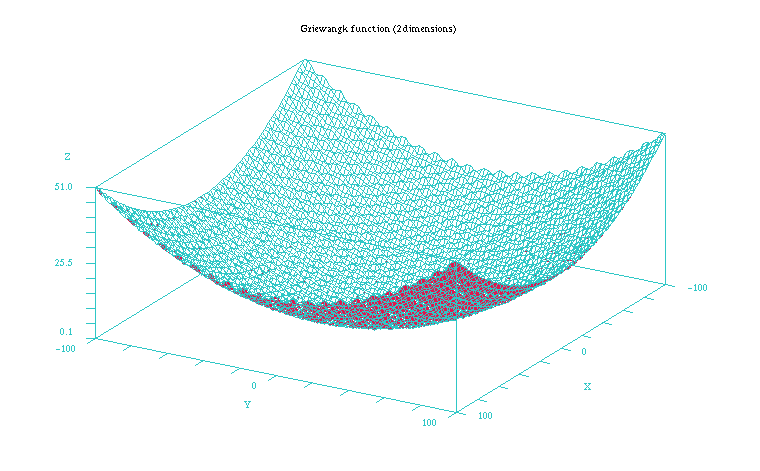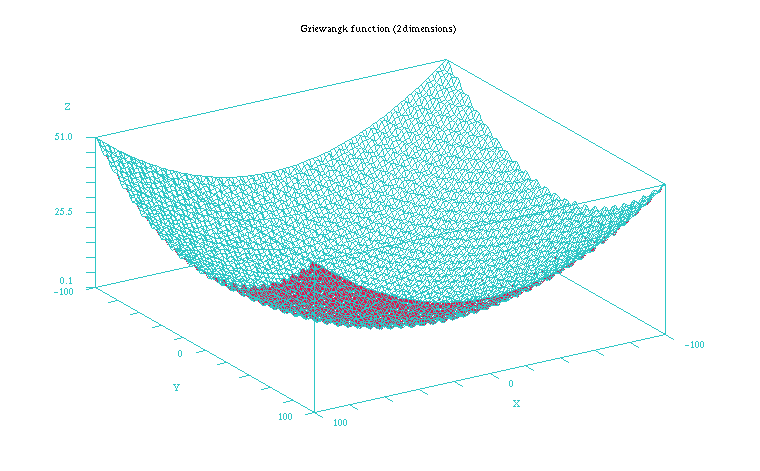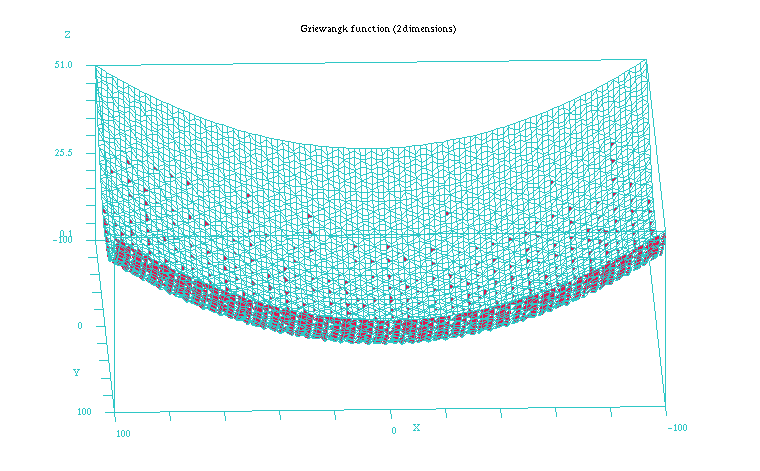
Figure 2: Plot of the Griewangk funtion on a closer 100x100 scale showing the local minima. Function computed with scilab.
| Dimension | Algorithm | Best fitness value | Number of evaluations |
| UMDAc | 6.0783E-02 ± 1.93E-02 | 301850 ± 0.0 | |
| MIMICc | 7.3994E-02 ± 2.86E-02 | 301850 ± 0.0 | |
| EGNABIC | 3.9271E-02 ± 2.43E-02 | 301850 ± 0.0 | |
| 10 | EGNABGe | 7.6389E-02 ± 2.93E-02 | 301850 ± 0.0 |
| EGNAee | 5.6840E-02 ± 3.82E-02 | 301850 ± 0.0 | |
| EMNAglobal | 5.1166E-02 ± 1.67E-02 | 301850 ± 0.0 | |
| EMNAa | 12.9407 ± 3.43 | 301850 ± 0.0 | |
| ES | 3.496E-02 ± 1.81E-02 | 25000 ± 1699.7 | |
| UMDAc | 8.9869E-06 ± 9.36E-07 | 177912 ± 942.3 | |
| MIMICc | 9.0557E-06 ± 8.82E-07 | 177912 ± 942.3 | |
| EGNABIC | 1.7075E-04 ± 6.78E-05 | 250475 ± 18658.5 | |
| 50 | EGNABGe | 8.6503E-06 ± 7.71E-07 | 173514.2 ± 1264.3 |
| EGNAee | 9.1834E-06 ± 5.91E-07 | 175313.3 ± 965.6 | |
| EMNAglobal | 8.7673E-06 ± 1.03E-06 | 216292 ± 842.8 | |
| ES | 1.479E-03 ± 3.12E-03 | 109000 ± 13703.2 |
Table 1: Mean values of experimental results after 10 executions for the problem Griewangk with a dimension of 10 and 50 (optimum fitness value = 0).
4.3 The evolution of the search
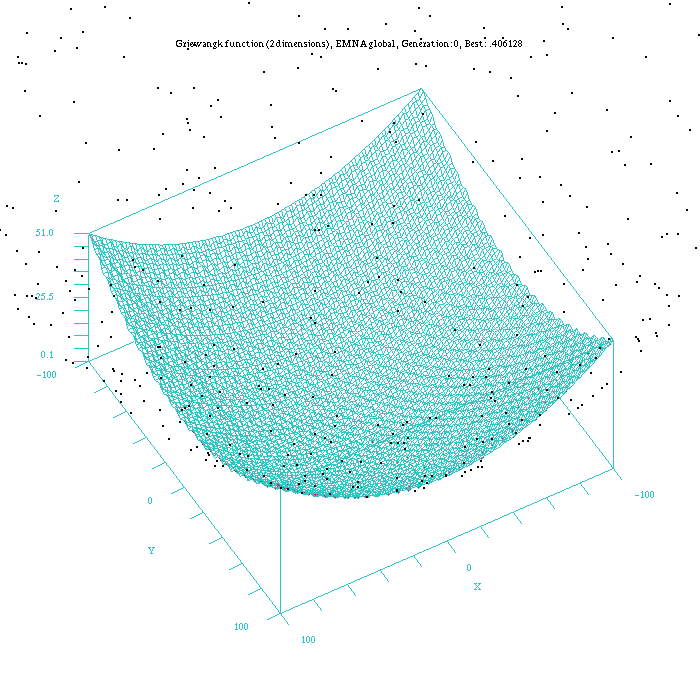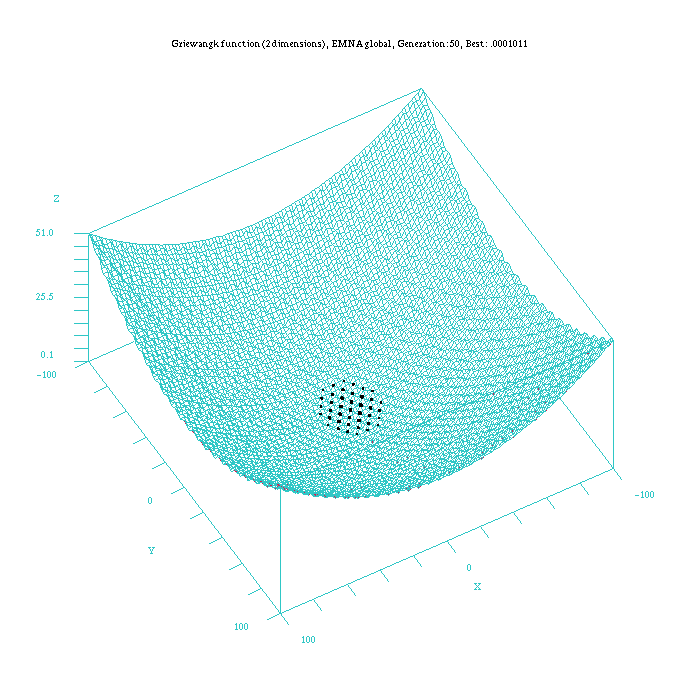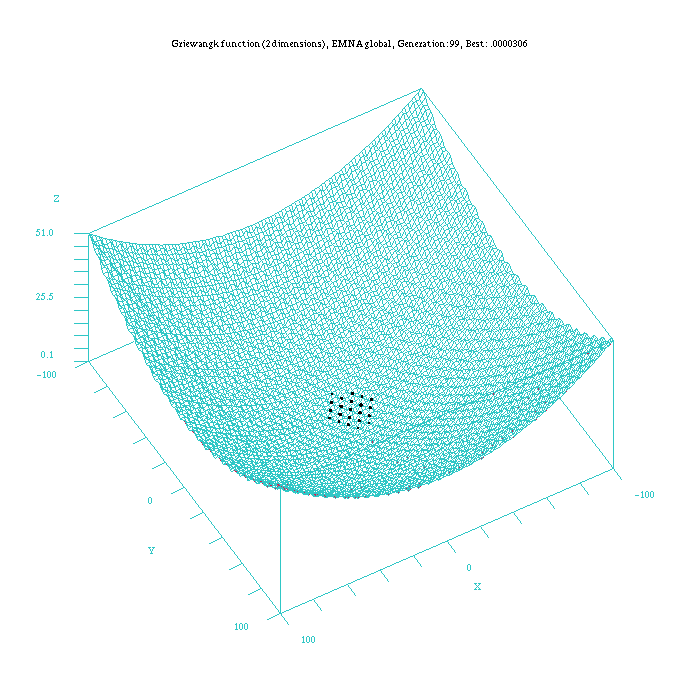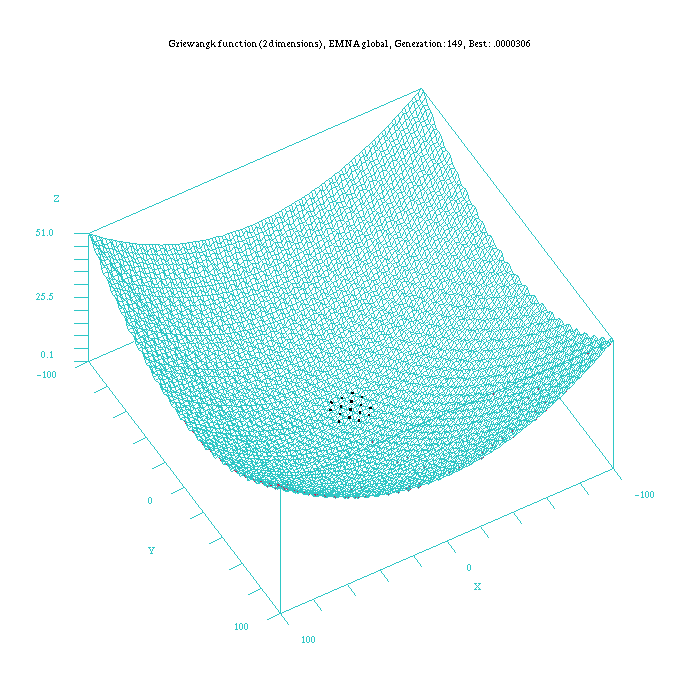
As an example to illustrate the difference in the way of reaching the final result for all the continuous EDAs, Figure 3, 4, 5, 6, and 7, show which is the behaviour of the UMDAc, MIMICc, Edge Exclusion, EMNA-global and EMNA-adaptative algorithms for the Griewangtk problem with a dimension of 2.
Figure 3: Plot of the UMDAc approach for the Griewangk funtion. Function computed with scilab.
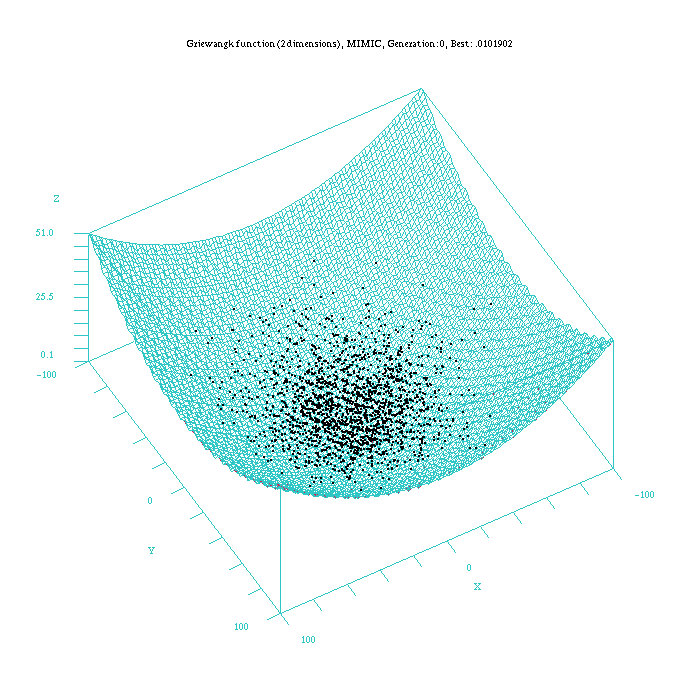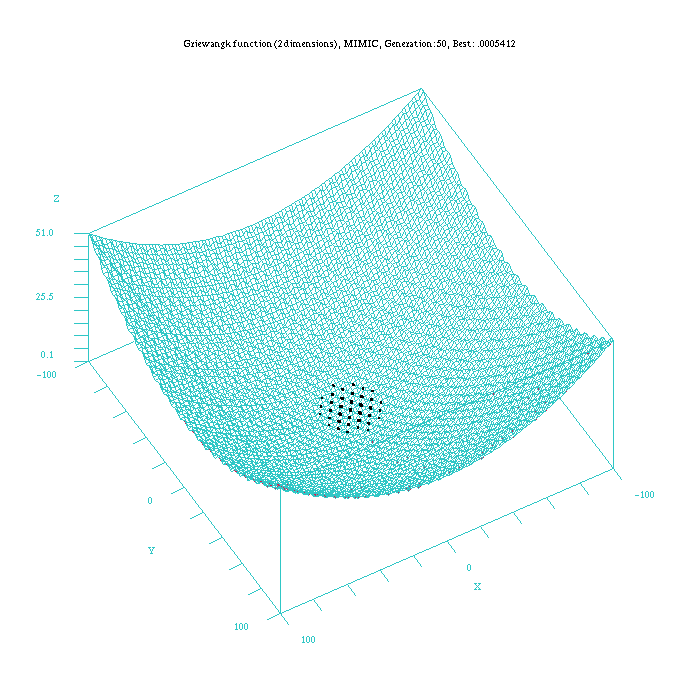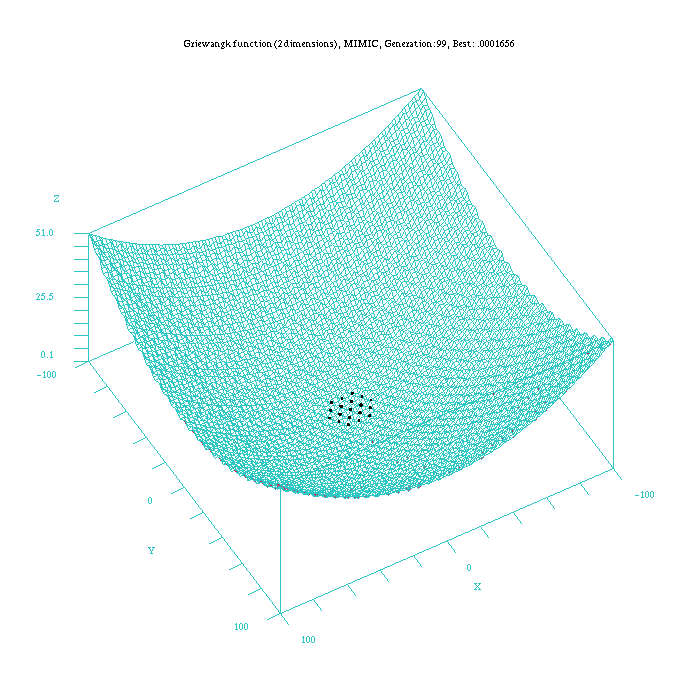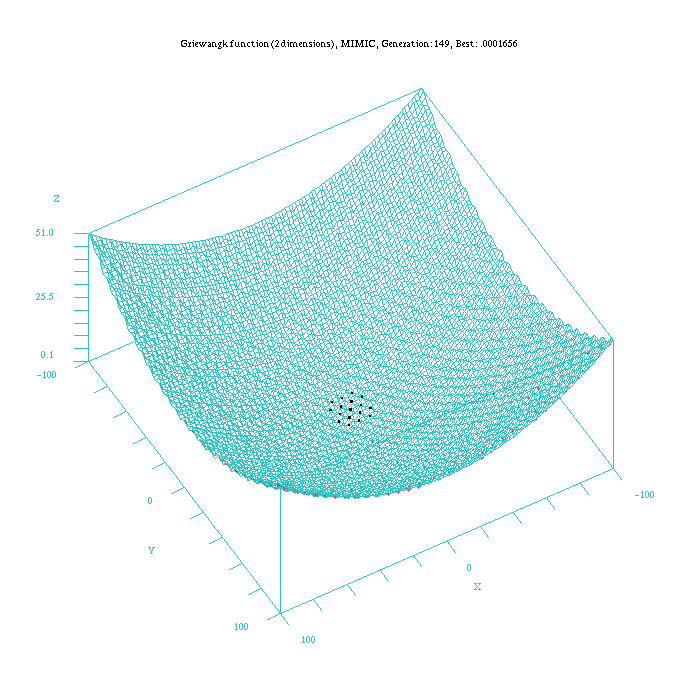
Figure 4: Plot of the MIMICc approach for the Griewangk funtion. Function computed with scilab.
Figure 5: Plot of the Edge Exclusion approach for the Griewangk funtion. Function computed with scilab.
Figure 6: Plot of the EMNA-global approach for the Griewangk funtion. Function computed with scilab.
Figure 7: Plot of the EMNA-adaptative for the Griewangk funtion. Function computed with scilab.
5 Conclusions
At the light of the results obtained in the fitness values, we can conclude the following: generally speaking, for less dimensions EMNAglobal, and EGNA type algorithms perform better that the rest, but when increasing the dimensions some of the algorithms show a poorer performance as a result of the higher complexity to take into account (e.g. the case of EGNABIC). The EMNAa algorithm showed a very poor behaviour for all these optimization problems, and its additional computation effort made impossible to apply it to the 50 dimension cases.
An important aspect to take into account is that the EMNAglobal algorithm appears to be the method that more quickly approaches to the best results, although these results are not always the optima. Nevertheless, once this algorithm is nearby the optimum solution it requires more time than algorithms as EGNABGe or EGNAee to satisfy the ending criterion.
Depending on the problem the ES method showed better results than the continuous EDAs, but when the type of problem to optimize presents many local minima and maxima, continuous EDAs show a more appropriated behaviour. The main drawback for continuous EDAs in general is the computation time they require, but for some problems the results that can be obtained with them are not comparable to methods in the ES category.
References
-
- [ Balas and Vazacopoulos1998]
- E. Balas and A. Vazacopoulos. Guided local search with
shifting bottlenck for job shop scheduling. Management Science,
44:262-275, 1998.
-
- [ Baluja and Davies1998]
- S. Baluja and S. Davies. Fast probabilistic modeling for
combinatorial optimization. In Proceedings of AAAI-98 Conference,
1998.
-
- [ Bengoetxea et al. 2000]
- E. Bengoetxea, P. Larrañaga, I. Bloch, A. Perchant,
and C. Boeres. Inexact graph matching using learning and simulation
of Bayesian networks. An empirical comparison between different approaches
with synthetic data. In Proceedings of CaNew workshop, ECAI 2000
Conference, ECCAI, Berlin, aug 2000.
- [ Bengoetxea et al. 2001]
- E. Bengoetxea, P. Larrañaga, I. Bloch, and
A. Perchant. Solving graph matching with EDAs using a permutation-based
representation. In P. Larrañaga and J. A. Lozano, editors,
Estimation of Distribution Algorithms: A New Tool for Evolutionary
Computation. Kluwer Academic Publishers, 2001.
- [ Blazewicz et al. 1996]
- J. Blazewicz, W. Domschke, and E. Pesch. The job
shop scheduling problem: Conventional and new solution techniques.
European Journal of Operational Research, 93:1-33, 1996.
- [ Buntine1991]
- W. Buntine. Theory refinement in Bayesian networks. In Proceedings
of the Seventh Conference on Uncertainty in Artificial Intelligence,
pages 52-60, 1991.
- [ De Bonet et al. 1997]
- J. S. De Bonet, C. L. Isbell, and P. Viola.
MIMIC: Finding optima by estimating probability densities. In Advances
in Neural Information Processing Systems, volume 9. M. Mozer,
M. Jordan and Th. Petsche eds., 1997.
- [ Etxeberria and Larrañaga1999]
- R. Etxeberria and P. Larrañaga. Global optimization
with Bayesian networks. In Special Session on Distributions and
Evolutionary Optimization, pages 332-339. II Symposium on Artificial
Intelligence, CIMAF99, 1999.
- [ Fisher and Thompson1963]
- H. Fisher and G.L. Thompson. Probabilistic learning of local
job-shop scheduling rules. In J.F. Muth and G.L. Thompson, editors,
Industrial Scheduling. Prentice-Hall, Englewood Cliffs, NJ,
1963.
- [ Henrion1988]
- M. Henrion. Propagating uncertainty in Bayesian networks by
probabilistic logic sampling. Uncertainty in Artificial Intelligence,
2:149-163, 1988. J.F. Lemmer and L.N. Kanal eds., North-Holland, Amsterdam.
- [ Larrañaga and Lozano2001]
- P. Larrañaga and J. A. Lozano. Estimation of
Distribution Algorithms: A New Tool for Evolutionary Computation.
Kluwer Academic Publishers, 2001.
- [ Larrañaga et al. 2000]
- P. Larrañaga, R. Etxeberria, J.A. Lozano, and J.M.
Peña. Optimization in continuous domains by learning and simulation
of Gaussian networks. In Proceedings of the Workshop in Optimization
by Building and using Probabilistic Models. A Workshop within the
2000 Genetic and Evolutionary Computation Conference, GECCO 2000,
pages 201-204, Las Vegas, Nevada, USA, 2000.
- [ Mühlenbein and Paaß1996]
- H. Mühlenbein and G. Paaß. From recombination
of genes to the estimation of distributions i. binary parameters.
In Lecture Notes in Computer Science 1411: Parallel Problem Solving
from Nature - PPSN IV, pages 178-187, 1996.
- [ Mühlenbein1998]
- H. Mühlenbein. The equation for response to selection
and its use for prediction. Evolutionary Computation, 5:303-346,
1998.
[ Törn and Zilinskas1989]- A. Törn and A. Zilinskas. Global Optimization. Lecture Notes in Computer Science 350. Springer Verlag, Berlin-Heidelberg.
Links on Estimation of Distribution Algorithms
The interested reader can find more links on EDAs here, inside the web page of the Intelligent Systems Group
File translated from TEX by TTH, version 2.89.
On 7 Mar 2001, 20:03. - A. Törn and A. Zilinskas. Global Optimization. Lecture Notes in Computer Science 350. Springer Verlag, Berlin-Heidelberg.